The new STED technique enables deep-tissue imaging, reveals subcellular dynamics of neurons.
А new microscopy technique that can acquire 3D super-resolution images of subcellular structures from about 100 microns deep inside biological tissue, including the brain, has been developed. That method could help reveal subtle changes that occur in neurons over time, during learning, or an as a result of the disease.
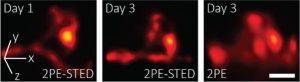
Researchers developed a 3D-2PE-STED system that can image dendritic spines deep inside the brain of a living mouse. Their system showed subtle changes that occurred between Day 1 and 3. These changes are hard to distinguish using two-photon microscopy alone. Photo Credits: Joerg Bewersdorf, Yale School of Medicine.
The new approach is an extension of stimulated emission depletion (STED) microscopy, which means that it achieves nanoscale resolution by overcoming the traditional diffraction limit of optical microscopes.
In Optica, The Optical Society’s (OSA) journal for high impact research, the researchers describe how they used their new STED microscope to image, in super-resolution, the 3D structure of dendritic spines deep inside the brain of a living mouse – dendric spines are tiny protrusions on the dendritic branches of neurons, which receive synaptic inputs from neighbouring neurons. They play a crucial role in neuronal activity.
The leader of their research team says that their microscope is the first instrument in the world to achieve 3D STED super-resolution deep inside a living animal.
Conventional STED microscopy is usually used to image cultured cell specimens. They use the technique to image thick tissue or living animals which is quite a challenge, especially when the super-resolution benefits of STED are extended to the third dimension for 3D-STED. This limitation happens due to the fact that thick and optically dense tissue prevents light from penetrating deeply and from focusing properly, thus impairing the super-resolution capabilities of the STED microscope.
To get over this, the researchers combined STED microscopy with two-photon excitation (2PE) and adaptive optics. They also added adaptive optics to their system. They claim:
“The use of adaptive optics corrects distortions to the shape of light, i.e., the optical aberrations, that arise when imaging in and through tissue.”
During imaging, the adaptive element modifies the light wavefront in the exact opposite way that the tissue in the specimen does and the aberrations from it. The aberrations from the adaptive element, therefore, cancel out the aberrations from the tissue, creating ideal imaging conditions that allow the STED super-resolution capabilities to be recovered in all three dimensions.
The researchers tested their 3D-2PE-STED technique by first imaging well-characterized structures in cultured cells on a coverslip, which resolved volumes more than 10 times smaller.
They made some tests, after which, they used their 3D-2PE-STED microscope to image the brain of a living mouse. They zoomed in on part of a dendrite and resolved the 3D structure of individual spines. They then imaged the same area two days later and showed that the spine structure had indeed changed during this time. No changes were observed, but the researchers will study this further. Velasco claims
“Dendritic spines are so small that without super-resolution it is difficult to visualize their exact 3D shape, let alone any changes to this shape over time, 3D-2PE-STED now provides the means to observe these changes and to do so not only in the superficial layers of the brain but also deeper inside, where more of the interesting connections happen.”








